Plot signal distribution histogram for accessibility threshold selection
Source:R/visualizations.R
plot_signal_histogram.RdVisualizes the distribution of BigWig signal across genomic regions to help identify the noise floor and select an appropriate z-score or raw signal threshold for calling accessible regions. The distribution is typically bimodal: a large peak at zero/low signal (background noise) and a smaller peak at higher signal (accessible regions).
Arguments
- bw_paths
named character vector of BigWig file paths.
- regions
GRanges object of regions to examine.
- n_bins
integer, number of histogram bins (default: 100).
- show_z_lines
numeric vector, z-score thresholds to show as vertical lines (default: c(1.0, 1.5, 2.0)).
- log_scale
logical, use log10(signal + 1) for x-axis (default: TRUE).
- cex_label
Numeric. Font size for threshold and annotation labels (default 0.7).
- cex_title
Numeric. Font size for histogram titles (default 1.0).
- hist_fill
character. Histogram bar fill color (default "#CCDDEE").
- hist_border
character. Histogram bar border color (default "#88AACC").
- density_col
character. Density overlay line color (default "#2C3E50").
- z_colors
character vector. Colors for z-score threshold lines (default: green, orange, red).
- suggest_col
character. Color for suggested threshold line (default "#B10DC9").
Value
Invisible list of per-sample signal statistics, including mean, sd, quantiles, and suggested thresholds.
Examples
bw_path <- make_example_bigwig()
regions <- GenomicRanges::GRanges(
"chr1", IRanges::IRanges(
start = c(1000L, 5000L, 10000L, 20000L, 50000L),
end = c(2000L, 6000L, 11000L, 21000L, 51000L)
)
)
stats <- plot_signal_histogram(c(Sample = bw_path), regions)
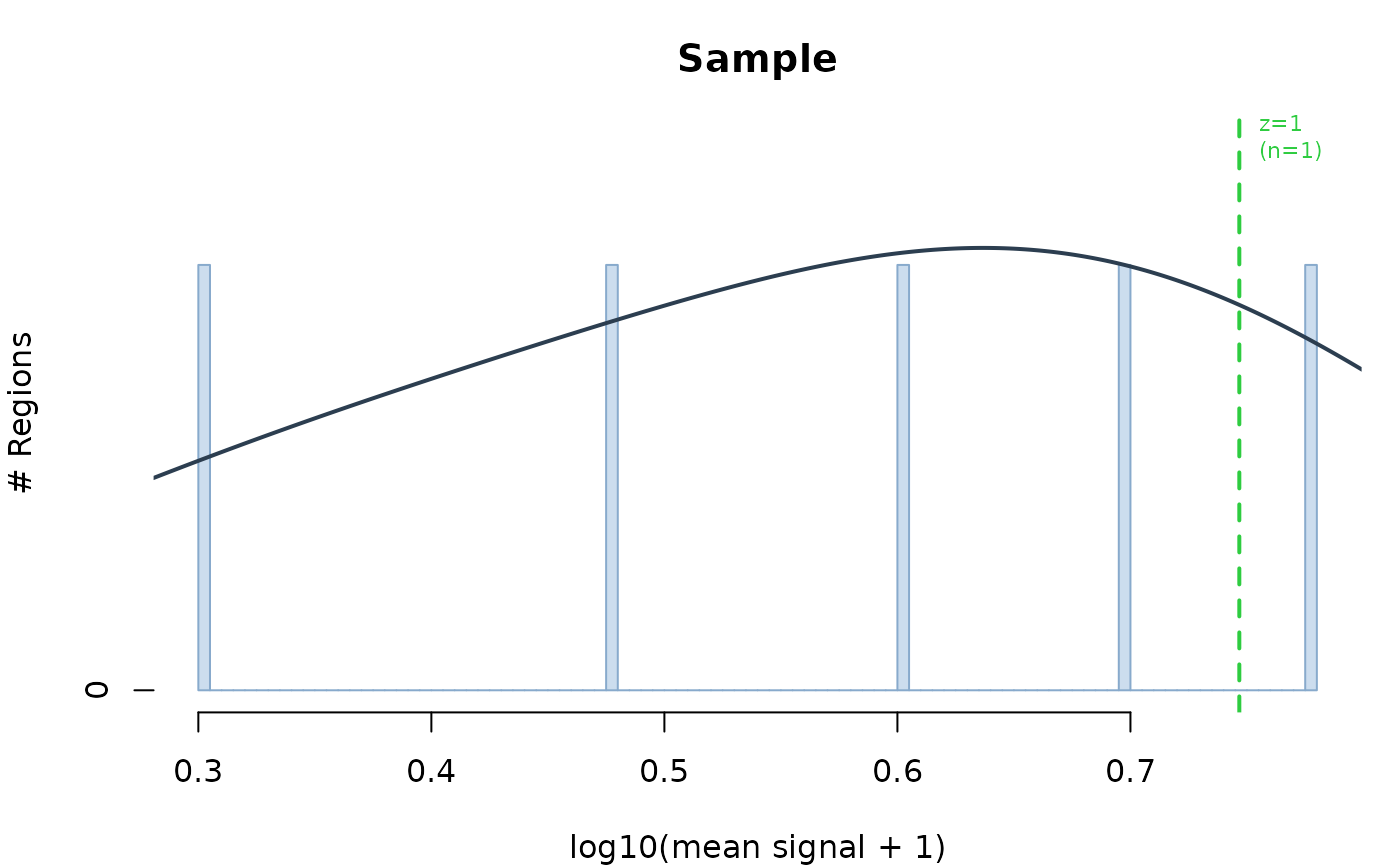 file.remove(bw_path)
#> [1] TRUE
file.remove(bw_path)
#> [1] TRUE